Research Progress
Information Manifolds of Cell Phenotype Transitions
How cells transition from one state to another—known as cell phenotype transition (CPT)—has long been a central question in fields such as developmental biology and disease pathogenesis (e.g., cancer metastasis). In recent years, the explosion of single-cell RNA sequencing technology has provided massive amounts of data, yet extracting physically and geometrically meaningful dynamic laws from high-dimensional gene expression data remains a formidable challenge for scientists.
Although cells contain tens of thousands of genes, the actual dynamics of cell phenotype transitions are confined to a low-dimensional manifold due to the stringent constraints imposed by underlying gene regulatory networks. However, existing single-cell data analyses largely rely on dimensionality reduction and visualization tools such as t-SNE, UMAP, and diffusion maps. While these methods attempt to preserve topological structures when projecting high-dimensional data into two or three dimensions, they distort the true intrinsic geometric properties of the manifold. Such distortions in distances and neighborhoods can lead to situations where the cell developmental trajectories or differentiation paths observed by researchers on 2D plots may completely deviate from the actual dynamic processes in high-dimensional space.
In response to this challenge, this study proposes a novel theoretical framework called SCIM (Single Cell Information Manifolds). For the first time, the study introduces information geometry into the analysis of single-cell evolutionary dynamics. By employing a Gaussian embedding neural network to transform single-cell gene expression profiles into multivariate Gaussian distributions, the Fisher information metric is naturally defined in probability space, successfully establishing a Riemannian metric for the low-dimensional manifold underlying cell phenotype transitions. This approach overcomes the limitation of traditional dimensionality reduction methods, which are primarily used for visualization, and enables, for the first time, quantitative monitoring of the intrinsic geometric properties of cell phenotype transition manifolds.
Using this method, we discovered that cells located in low-curvature regions of the manifold correspond to critical stages of cell fate determination and highly unstable transitional-state cells. SCIM is the first to integrate RNA velocity with the Fisher information metric, calculating an "information velocity" for each cell. As a measure of how rapidly a distribution changes, information velocity can characterize the speed of cell state transitions. The study found that cells in low-curvature regions of the manifold exhibit high information velocity, reflecting the dynamic properties of transitional-state cells, and revealed that cell phenotype transitions resemble the principle in general relativity where "spacetime tells matter how to move"—across different systems, the dynamics of cells are guided by the geometric structure of the manifold.
The study of critical points in complex systems remains an active area of research. Information geometry and Fisher information offer a powerful approach for characterizing phase transition processes. Because this method does not require a predefined Hamiltonian or ensemble function, it can be readily applied to non-equilibrium systems. Therefore, the SCIM model not only provides a theoretical framework for analyzing single-cell phenotype transitions but also holds promise for uncovering deep principles underlying transition processes in a wide range of complex systems.
The related research findings, published in Cell Systems under the title "Geometric Quantification of Cell Phenotype Transition Manifolds with Information Geometry," offer a novel mathematical and physical perspective for quantifying non-equilibrium dynamics in complex biological systems.
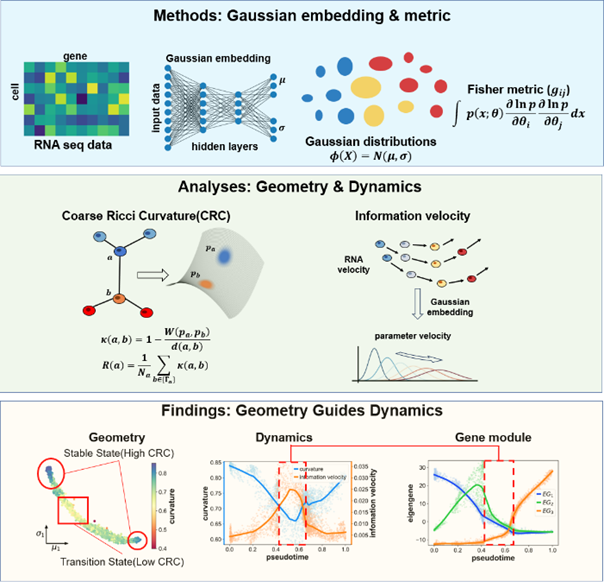
▲ Graph abstract of SCIM
Dr. Weikang Wang, Associate Professor at the Institute of Theoretical Physics, Chinese Academy of Sciences, and Professor Lei Zhang, from the Beijing International Center for Mathematical Research, the School of Mathematical Sciences, the Center for Quantitative Biology, and the International Center for Machine Learning at Peking University, serve as the co-corresponding authors of this paper. Miao Huang, a Ph.D. student at the Institute of Theoretical Physics, Chinese Academy of Sciences, and Yuxuan Wang, a master’s student at the Center for Data Science, Peking University, are the co-first authors. The paper’s collaborators also include Professor Haijun Zhou and Ph.D. student He Xiao from the Institute of Theoretical Physics, Chinese Academy of Sciences, as well as Junda Ying, a Ph.D. student at the School of Mathematical Sciences, Peking University. This research was supported by multiple funding sources, including the National Natural Science Foundation of China, the Special Fund for Theoretical Physics, the National Key Research and Development Program of China, and the Beijing Natural Science Foundation.

Article link: https://authors.elsevier.com/a/1mqz88YyDfyNHF
Contributor:Wei-Kang Wang
